Absolute proof: The Gp-120 sequences prove beyond all doubt that "COVID-19" was man-made
The "missing link" was there in Pradhan's paper all along, we just needed to ask the right question: "where are the genome sequences for the Gp-120 inserts"
Following on from December’s article “How to BLAST your way to the truth about the origins of COVID-19” where I explained how the four “inserts” described in Pradhan’s Feb 2020 paper showed definitively that the SARS-CoV-2 genome had to have been man-made, there has been considerable interest and a lot of push-back from those whose reputations are on the line.
From what I can make out the main lines of rebuttal were one of these:
“You are an anonymous troll” (anonymous - yes, a troll - only to those people)
“those sequences exist in bacteria and it doesn’t matter that viruses can’t combine with bacterial sequences” (OK, no problem, let’s just turn virology on its head and go with your new dogma that you’ve just plucked out of thin air)
“that 19nt sequence wasn’t actually claimed in the Moderna patent” (true, but it was still a Moderna sequence listed in the patent - they just didn’t want to patent all 30,000 of them because it would cost a fortune so they tricked Genbank into accepting them as patented sequences for free)
I’m not sure there were any others but feel free to correct me in the comments. Anyway, not bad I guess from an anonymous troll. So, now we have got the shills and detractors to use up their ammunition I am going to finish this argument, for once and for all, using the same BLAST techniques I showed you last time (so that you can verify for yourself that this virus was man-made). This time we’re going to dig into the genome sequences behind Pradhan’s “Inserts 1-3” that nobody has talked about. Until now.
TLDR: In order to get the 3 inserts of Gp-120 to exist in SARS-Cov-2, the genomic sequences that coded for them had to have got there by recombination from another organism or in a lab. Because they don’t exist anywhere in nature it is not possible to have come from another organism. Ecohealth's proposal in 2018 perfectly described those inserts. Until now we have only discussed the peptide (i.e. amino acid) construction of the 4 sequences referenced in Pradhan’s paper that showed that the main differences between the spike protein of SARS-CoV-2 and any other previous coronavirus known were contained in these “Inserts”. Here they are:
We’re not going to discuss insert 4 for now because we discussed it in part 1, but safe to say that Insert 4 is a quite unique and interesting sequences because it not only codes for a very useful Furin Cleavage Site (FCS) - which increases the virulence of the SARS-like virus, but it also has homology to the HIV-Gag protein as you can see. It also contains the unique Moderna nucleotide sequence we discussed in part 1 and which has now been published - after a long wait - by Balamurali Ambati here.
We don’t need to discuss it in this article because there is enough damning evidence in the other inserts to lay this to rest. Just to put this into perspective I’m going to show you where these inserts are in the actual spike protein molecule (and, of course, in the Pfizer & Moderna “vaccine” spike protein, which is identical in this area of the molecule).
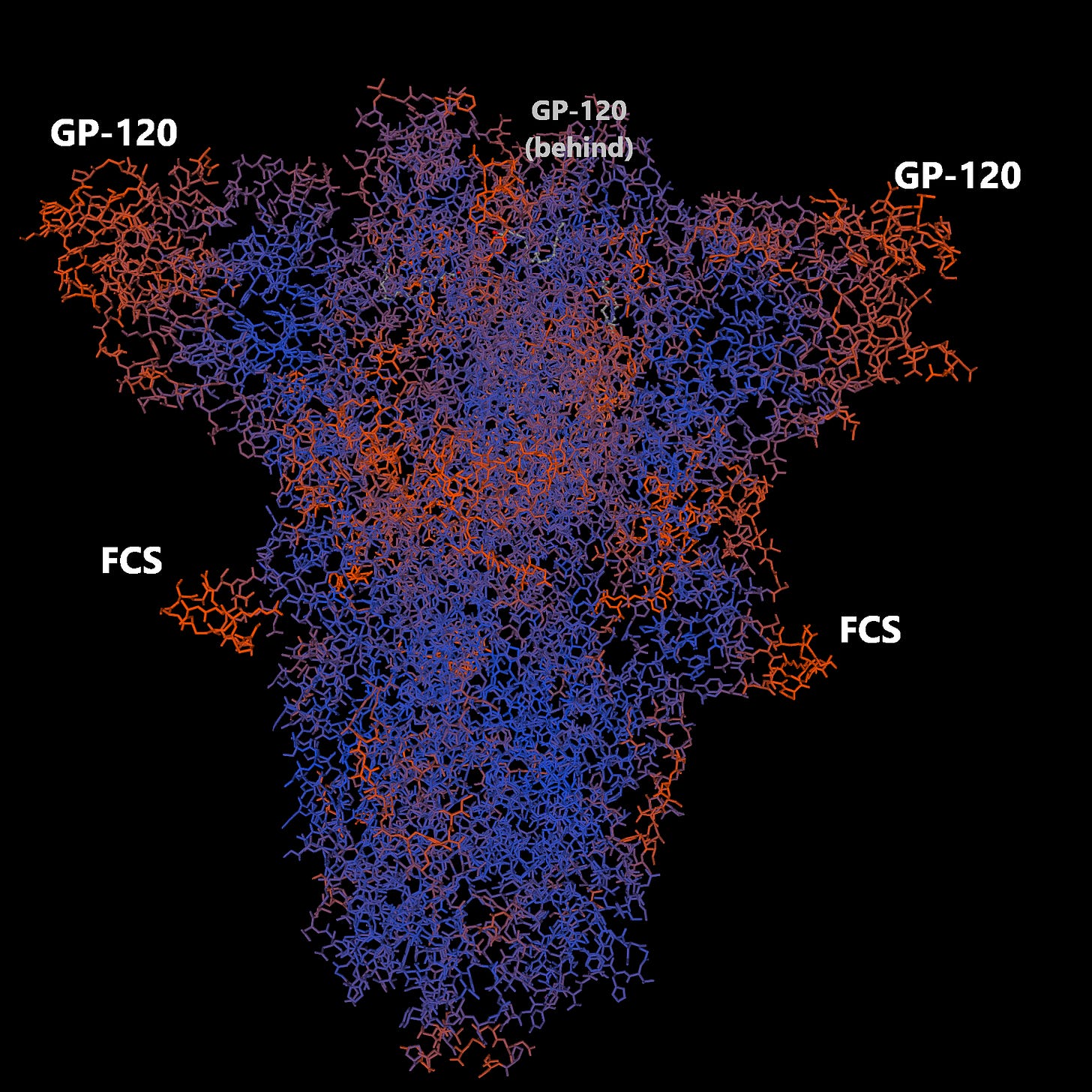
What is shown is a 3D model of the spike protein trimer (3 spikes held together due to their conformation and amino acid makeup) so you can see that it makes a bit of a pyramid shape. The right hand picture shows the trimer looking down from the top of the left hand picture. Imagine flowers in a vase. The picture on the left is looking side-on and the picture on the right would be looking top-down
Here’s another view from the SWISSMODEL viewer. In our flower pot analogy the third flower is at the back in the middle. You will notice that the FCS inserts (insert 4) are in a strategic location. In the flowerpot analogy these are the handles of the flower-pot. From the top you wouldn’t see them very well. But the Gp-120 inserts are in a really strategic location. They are the flowers that every one can see, because they are sticking out. In a viral sense they are the most geographically likely to bind to something (i.e. a cell receptor) so they are really strategic.
In fact, most of what the spike protein does now depends on these fragments binding. If you still aren’t with me, imagine you are a cat owner and the flowers are lilies (lilies are poisonous to cats). You would really need to keep your flower pot out of the way of your beloved feline to avoid those beautiful but poisonous lily heads killing your cat.
OK so hopefully the analogies are now sufficient to understand that we are dealing with viral protein inserts at a strategic location on a viral spike protein. But I need to bring this message home with another diagram:
The diagram shows something that is so coincidental that it already impossible that this has happened by chance. That is, that each of the 3 Gp-120 inserts described in Pradhan’s paper are located at the outermost strategic points in the whole viral spike. Those pangolins - to have managed to get the most important viral peptides in human immunology to have situated themselves in exactly the locations needed to infect human T-cells, in a bat cave (without there being any humans in the bat cave to practice on) - are clever, I must admit. OK, I’m being sarcastic. It’s not possible and it doesn’t matter how many pangolins and bats got together in a bat-o-lin orgy. As I said before - not gonna happen.
But, that’s not all. You see, all the shills and detractors and people like “Fast Eddie” Holmes (who you can read about here) are going to tell you that these peptides are still too small to be able to be used as the ultimate smoking gun in the case for lab origins. Well, that’s true but irrelevant - because there are two other aspects that make this impossible to be natural.
Aspect #1: The function of these inserts - that didn’t exist before 2019 in any coronavirus - were clearly stated as the intention of the DARPA DEFUSE proposal
In case you haven’t seen it, this is the document (archived here). You can see the involved players - Peter Daszak and Ecohealth Alliance.
If you look through the proposal you probably won’t make much of it because it looks like they were actually trying to look at “defusing” the threat of coronaviruses. The only problem is that they weren’t doing that at all. This is what they said they were going to do. In their words (the yellow highlights are the most important bits):
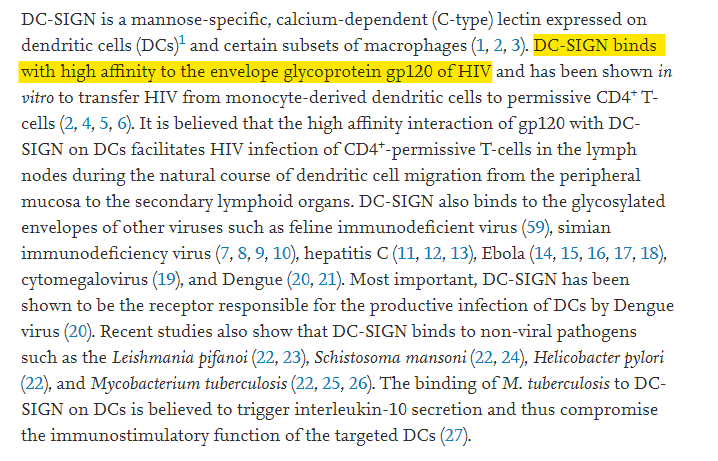
So they were going to introduce furin cleavage sites and sites binding DC-SIGN into coronaviruses (what could possibly go wrong?). Well, we know about the furin cleavage site already. What the hell is DC-SIGN, and what is needed to bind it.
Well, you might think this is a coincidence but….
That’s just one paper, there are many on DC-SIGN <=> Gp120 binding. Essentially it’s part of the mechanism that HIV-1 uses to bypass the immune system and gain entry to the very cells that keep your immune system working (T-cells).
This means that the Ecohealth proposal in 2018 described exactly the two changes made to existing SARS-coronaviruses that resulted in SARS-CoV-2 in 2019. Billions of years of random evolutionary change (that could never have been possible because there were no humans living in the bat cave for the evolutionary changes to promulgate) happened in 1 year. Then the authors of the proposal and all of their connected chums went into overdrive to dispel any talk of what you can now see is completely obvious. (If you haven’t seen Chris Martenson’s excellent review of the people involved and their nefarious meeting you should).
Had enough yet? Well, I haven’t because this is the final piece that proves irrefutably (as if the above wasn’t enough) that this was man-made.
Aspect #2: The genome sequences coding those Gp-120 loops don’t exist
What the hell does that mean? Well, in order for the pangolins to get some viruses together and accidentally (naturally) create a new one with their clumsy claws, they need another organism that has some genomic sequences (GATTACA… etc) that can transfer to the old virus to make a new one. It’s called recombination. I mean, sure, the old virus can undergo some mutations (GATTTCA…) but these are evolutionarily very slow and can result in deletions and changes. In order to get inserts for these to happen by chance is so rare that it would take millions of years to develop functional inserts by chance so you really need a genome donor. Essentially you need to make a chimaera (RNA donated from another organism to the first organism to make a new organism). In other words, for inserts of this size that are functional the genome sequences coding for them must exist somewhere (because they have to be donated to form the chimaeric RNA)
So let’s find out shall we? All we have to do is plug the genome sequences into BLAST and see if those sequences exist in any other virus (or organism that can recombine with a virus). On the latter note, it’s important to be clear that bacterial DNA can’t recombine with RNA viruses in nature.
But let’s just accept that for a minute. We have 3 Gp-120 (HIV) peptide inserts to blast: TNGTKR, HKNNKS and RSYLTPGDSSSG. Two of them are 6-mers (requiring 18 nucleotides to encode, 3 for each amino acid) and the last is a 12-mer (requiring 36nt). Remember that recombination between viruses occurs at the genomic level (the nucleotides e.g. GATTACA) not the peptide level. In other words, to show a recombination event has occurred to insert TNGTKR we need to find the genome sequence that encoded TNGTKR and see if it exists in another virus. So, we need the genome sequences for these peptides and here they are:
TNGTKR is encoded by acc aat ggt act aag agg
HKNNKS is encoded by cac aaa aac aac aaa agt
RSYLTPGDSSSG is encoded by aga agt tat ttg act cct ggt gat tct tct tca ggt
Immediately you can see that the longer one will be the most interesting (because the probability of a coding occurring by change is approximately 1 in 4^n where n in the number of nucleotides. So for a 36-mer this is one in 4-billion-trillion or so). But for the smaller ones we will need to restrict to viruses again. Let’s do it, let’s get back to BLAST. You’ll need to be quite restrictive for an 18-mer so we’ll ask for 100% sequence homology, just viruses and nothing from 2020, 2021 or 2022 (because most of the hits will be sequences occurring after SARS-CoV-2 was “discovered”)
Here’s the first one for which I’ll show the BLASTn screen…
and the output, which I think speaks for itself…
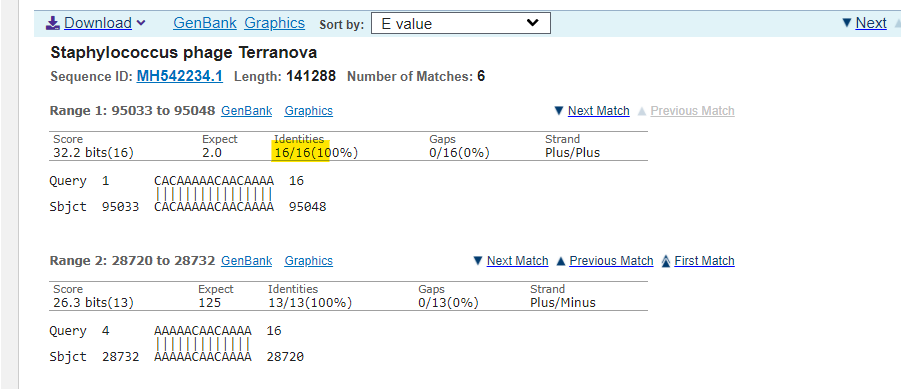
The output for Insert 2 is more interesting and busy, so requires a filter for percent identity (=100%) and Query coverage (=100%) and this is the output:
Pretty much all of those are phages, which can be used in the lab to transfer genomes to bacteria. But although those all appear to be 100% alignment sometimes BLAST does strange things and in fact none of the list matches all 18 nucleotides. That is, there was no 18/18 match to this sequence in viruses.
So, of our two sequences so far we have two that don’t match any viruses fully (18/18nt) and the closest match is to HIV-1 and a bacterial phage. Hmmmm….
Now the last one. Because it’s a longer sequence we can be less restrictive with our search, just excluding “SARS-Cov-2” and “Synthetic constructs”. We can use Megablast for highly similar sequences. This is the result:
And there we have it. 3 gene sequences.
None of them exist in viruses.
One of them doesn’t exist in nature at all.
Yet they appear in a virus that arose from nowhere, 1 year after Ecohealth said they were going to make one that did exactly what they proposed. And they produce peptides that arise exactly on the binding sites of the virus.
As I said, those Pangolins - they’re smart I tell ya’.
[NOTE: for a deeper diver into the EcoHealth and Fauci cover-up relating to this please make Charles Rixey’s blog your follow - he has done some great work. ]


















Is there any virologist,strong enough ( and old enough not to care about fall out 🤣😂)that could present your brilliant work? Whilst preserving your anonymity?
I am 70+ and trying to educate my peers to what is happening.Most people with mortgages and children can’t go public. What about Project Veritas. Thank you for searching for the truth.
I will have to read this several more time to be able to explain it to someone else.O level biology doesn’t quite cover it.😊
Thank you for looking these up!
And indeed the Sars-Cov-2 virus can infect T cells via one mechanism and monocytes through the other.
This is the key feature of how repeat infections of the vaxxed (caused by Original Antigenic Sin and vaccine immune suppression), will kill off above mentioned immune cells and will create an analogue of AIDS in the unfortunate victims. These inserts are there for a reason.
Sars-Cov-2 is already "highly pathogenic", no need to wait for it to deattenuate further although it could become even more pathogenic.
I call the result of this "Chronic Covid" (not the same thing as "endemic covid" or "long covid"). Chronic Covid is a covid infection in person who no longer can clear the virus.
https://igorchudov.substack.com/p/aids-like-chronic-covid-is-taking